Introduction
(Article introduction authored by Conquest Editorial Team)
In kidney transplantation, day-zero biopsies are used to assess organ quality and discriminate between donor-inherited lesions and those acquired post-transplantation.
However, many centers do not perform such biopsies since they are invasive, costly and may delay the transplant procedure.
We aim to generate a non-invasive virtual biopsy system using routinely collected donor parameters. Using 14,032 day-zero kidney biopsies from 17 international centers, we develop a virtual biopsy system.
11 basic donor parameters are used to predict four Banff kidney lesions: arteriosclerosis, arteriolar hyalinosis, interstitial fibrosis and tubular atrophy, and the percentage of renal sclerotic glomeruli.
Six machine learning models are aggregated into an ensemble model. The virtual biopsy system shows good performance in the internal and external validation sets. We confirm the generalizability of the system in various scenarios.
This system could assist physicians in assessing organ quality, optimizing allograft allocation together with discriminating between donor derived and acquired lesions post-transplantation.
Methods
The study encompassed a diverse population of adult kidney donors for transplantation, spanning from 2000 to 2021, with pre-transplantation kidney biopsies performed as part of standard care. Involving 15 centers across seven countries for derivation and two external validation institutions, a total of 15,121 kidney biopsies were evaluated, with 14,032 included for final analysis after excluding inadequate samples.
Data were anonymized and collected in compliance with ethical standards, and the study protocol was approved by the Paris Transplant Group’s Institutional Review Board. Expert kidney pathologists graded biopsy lesions according to the international Banff classification system.
Analyses were conducted using machine learning algorithms and statistical methods, with rigorous internal and external validation to assess model performance. Imputation techniques addressed missing data, while a sensitivity analysis evaluated the predictive capability of the Kidney Donor Profile Index.
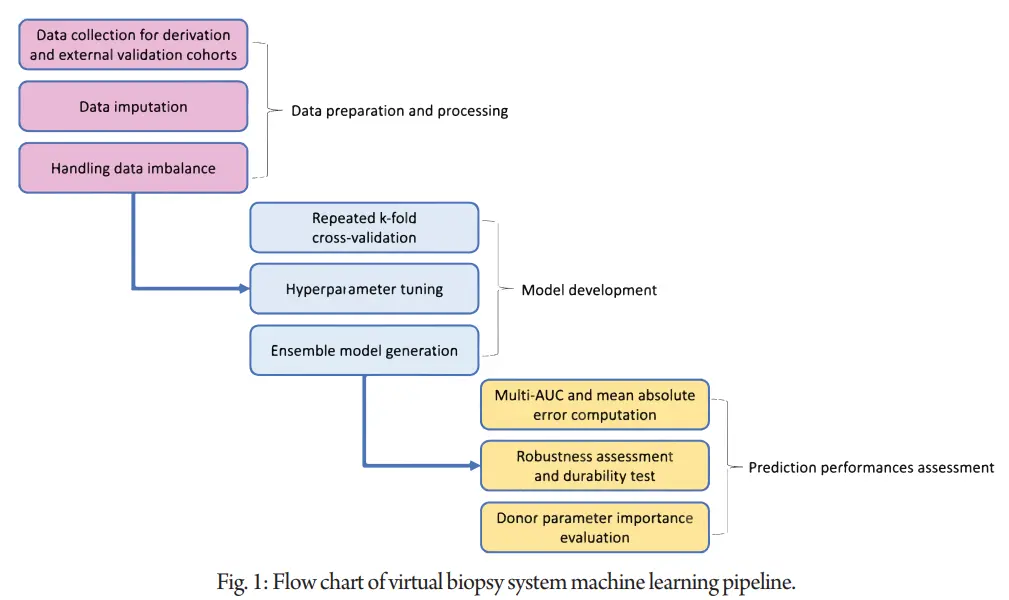
Consistency in biopsy evaluation was examined through inter-pathologist reassessment. Overall, the study employed robust methodologies to develop and validate a virtual biopsy system for predicting kidney histological lesions using donor parameters, holding potential for improving pre-transplant organ assessment and patient outcomes in kidney transplantation. Figure 1 summarizes the process of generating and validating machine learning models.
Results
Kidney virtual biopsy system development
The population cohort was imputed separately by derivation and external cohorts then pre-processed. We tuned and generated the best performing models for predicting the lesion scores, based on the donor parameters. Then, the ensemble model that groups these models together was generated. For each biopsy lesion score, we selected the ensemble models as a virtual biopsy system.
Kidney virtual biopsy system development
We examined the importance of the 11 donor parameters used for the virtual biopsy system development by averaging the importance produced by the models.

Model prediction performance on derivation cohort
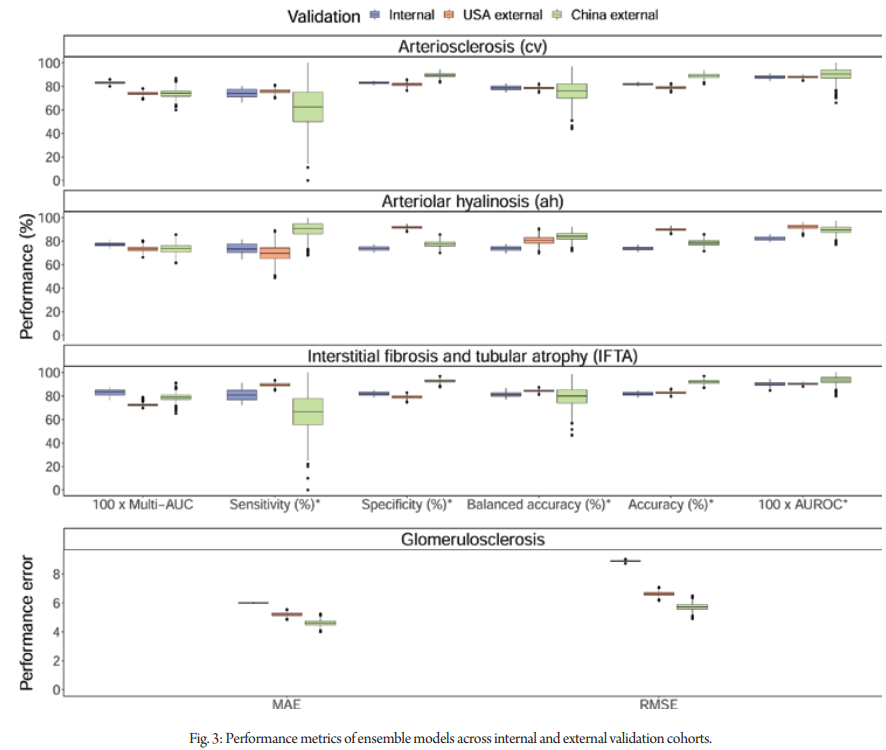
The ensemble models showed discrimination performance during cross-validation with the multi-area under the curves (multi-AUC) of 0.833 (SD 0.013), 0.773 (0.020), 0.830 (0.027) for cv, ah, and IFTA lesions, respectively.
Additionally, the ensemble models achieved area under the receiver operating characteristic curves (AUROC) of 0.880 (0.016), 0.823 (0.019), and 0.900 (0.023) for cv, ah, and IFTA lesions, respectively. Ensemble models’ cut-offs were calibrated to maximize Youden’s J statistic. With the calibrated cut-offs of 0.582 for cv, 0.596 for ah, and 0.637 for IFTA, balanced accuracies (mean of sensitivity and specificity) were 0.786 (0.021) for cv, 0.736 (0.021) for ah, and 0.813 (0.024) for IFTA.
For the glomerulosclerosis lesion, the mean absolute error (MAE) was 5.999 (0.032) and the root mean square error (RMSE) was 8.888 (0.059). The ensemble models and random forest models showed comparative performance.
External validation of the virtual biopsy system
We included 1630 day-zero biopsies from the USA and China for external validation. The median percentage of glomerulosclerosis was 2.1% (IQR 0.0-12.5). The distribution of cv lesion scores was 27.9% (None), 33.9% (Mild), 36.3% (Moderate), and 1.9% (Severe). For ah lesion scores, the distribution was 53.8%, 38.4%, 6.4%, and 1.4%. IFTA scores distribution was 40.4%, 30.7%, 28.7%, and 0.2%. Most moderate or severe lesions (cv, ah, IFTA) were from deceased donors.
In the Columbia University cohort, ensemble models had multi-AUCs of 0.740 (cv), 0.733 (ah), and 0.723 (IFTA). The AUROCs were 0.880, 0.922, and 0.905, respectively. Balanced accuracies were 0.787, 0.808, and 0.843, respectively. For glomerulosclerosis, the MAE was 5.200 and RMSE was 6.630. In the Sun Yat-sen University cohort, ensemble models had multi-AUCs of 0.740 (cv), 0.736 (ah), and 0.798 (IFTA).
The AUROCs were 0.902, 0.895, and 0.935, respectively. Balanced accuracies were 0.760, 0.840, and 0.797, respectively. For glomerulosclerosis, the MAE was 4.608 and RMSE was 5.731.Figure 2 summarizes the performance of the ensemble models.
Validation of the virtual biopsy system in various scenarios
We confirmed the robustness of the virtual biopsy system in different subpopulations and clinical scenarios in the internal cross-validation, including (i) region (Europe, North America or Australia), (ii) donor ethnicity (African American, Caucasian, and Others [Hispanic, Asian, and Arabic]), (iii) donor criteria (extended criteria donors or standard criteria donors plus living donors), and (iv) biopsy type (preimplantation and postreperfusion). Overall, the system showed good performance in subpopulations. These analyses are depicted in Supplementary Table 12.
Pathologists’ biopsy findings reliability
We confirmed the inter-pathologist consistency in four expert nephropathologists from Necker hospital and Mayo clinic in evaluating the biopsy findings, with Fleiss Kappas of 0.68 (95% CI 0.63–0.73), 0.59 (0.53–0.65) and 0.51 (0.44–0.59), for cv, ah, and IFTA lesions respectively. The overall Fleiss Kappa for all lesions was 0.63 (0.60–0.66).
Performance of kidney donor profile index (KDPI) score
The derivation cohort included 4241 biopsies, and the external validation cohort comprised 1124 biopsies (920 from Columbia University medical center and 204 from Sun Yat-sen University). The mean KDPI was 53.43 (SD 29.49) in the derivation cohort and 63.24 (SD 26.63) in the external validation cohort.
Virtual biopsy system online application for physicians
Based on these results, we constructed a ready-to-use online application to offer physicians an open access to the virtual day-zero biopsy system (Supplementary Movie 1). The application allows physicians to enter a single patient’s data, to get (i) the personalized probabilities of belonging to each day-zero histological lesion score and (ii) the prediction visualization with radar chart.
Discussion
In this international, multicohort study, a virtual biopsy system was developed and validated using non-invasive donor parameters to predict kidney histological lesions. Four ensemble models were created, showing good discrimination, calibration, and generalizability across various countries and clinical scenarios.
The expansion of kidney donor pools, especially with older donors, has raised questions about organ quality assessment. However, the invasive and time-consuming nature of traditional biopsy procedures poses challenges, especially in time-sensitive transplantation settings. Existing studies lack a comprehensive virtual biopsy approach utilizing donor parameters.
The virtual biopsy system not only predicts lesion presence and severity but also aids in post-transplant lesion interpretation and patient monitoring. By leveraging high-quality data labeled by expert pathologists, it addresses issues of variability in biopsy interpretation.
Moreover, the system has potential implications for optimizing organ allocation, reducing cold ischemia time, improving patient stratification in clinical trials, and enhancing prognostic interpretation. An easy-to-use online application supports its practical applicability in real-world settings.
While the study has strengths, including robust validation and model assessment, limitations such as interobserver variability and biopsy technique heterogeneity are acknowledged. However, efforts were made to mitigate these limitations, ensuring the reliability and generalizability of the virtual biopsy system.
In conclusion, the machine learning-driven virtual biopsy system offers a promising solution for pre-transplant organ assessment, with implications for improving patient outcomes and resource utilization in kidney transplantation.
1. Mallory, T. B. Pathology. N. Engl. J. Med. 236, 438–443 (1947).
2. Barry, J. M. & Murray, J. E. The first human renal transplants. J. Urol. 176, 888–890 (2006).
3. Michon, L. et al. [An attempted kidney transplantation in man: medical and biological aspects]. Presse Med. 61, 1419–1423 (1953).
4. Gaber, L. W. et al. Glomerulosclerosis as a determinant of posttransplant function of older donor renal allografts. Transplantation 60, 334–339 (1995).
5. Naesens, M. Zero-time renal transplant biopsies: a comprehensive review. Transplantation 100, 1425–1439 (2016).
6. Mengel, M. et al. Protocol biopsies in renal transplantation: insights into patient management and pathogenesis. Am. J. Transpl. 7, 512–517 (2007).
7. Chauhan, A. et al. Using implantation biopsies as a surrogate to evaluate selection criteria for living kidney donors. Transplantation 96, 975–980 (2013).
8. Randhawa, P. Role of donor kidney biopsies in renal transplantation. Transplantation 71, 1361–1365 (2001).
9. Solez, K. et al. Banff 07 classification of renal allograft pathology: updates and future directions. Am. J. Transplant. 8, 753–760 (2008).
10. Sung, R. S. et al. Determinants of discard of expanded criteria donor kidneys: impact of biopsy and machine perfusion. Am. J. Transpl. 8, 783–792 (2008).
11. Mengel, M. & Sis, B. An appeal for zero-time biopsies in renal transplantation. Am. J. Transpl. 8, 2181–2182 (2008).
12. Springfield, D. S. & Rosenberg, A. Biopsy: complicated and risky. J. Bone Jt. Surg. Am. 78, 639–643 (1996).
13. Aubert, O. et al. Long term outcomes of transplantation using kidneys from expanded criteria donors: prospective, population based cohort study. BMJ 351, h3557 (2015).
14. Matas, A. J. et al. OPTN/SRTR 2013 Annual Data Report: kidney. Am. J. Transpl. 15, 1–34 (2015).
15. Jadlowiec, C. C. et al. Transplant outcomes using kidneys from high KDPI acute kidney injury donors. Clin. Transpl. 35, e14279 (2021).
16. Mancilla, E. et al. Time-zero renal biopsy in living kidney transplantation: a valuable opportunity to correlate predonation clinical data with histological abnormalities. Transplantation 86, 1684–1688 (2008).
17. Bora, A. et al. Predicting the risk of developing diabetic retinopathy using deep learning. Lancet Digit Health 3, e10–e19 (2021).
18. Miles, J., Turner, J., Jacques, R., Williams, J. & Mason, S. Using machine-learning risk prediction models to triage the acuity of undifferentiated patients entering the emergency care system: a systematic review. Diagn. Progn. Res 4, 16 (2020).
19. Sufriyana, H. et al. Comparison of multivariable logistic regression and other machine learning algorithms for prognostic prediction studies in pregnancy care: systematic review and meta-analysis. JMIR Med. Inf. 8, e16503 (2020).
20. Huang, P. et al. Prediction of lung cancer risk at follow-up screening with low-dose CT: a training and validation study of a deep learning method. Lancet Digit Health 1, e353–e362 (2019).
21. Roufosse, C. et al. A 2018 reference guide to the banff classification of renal allograft pathology. Transplantation 102, 1795–1814 (2018).
22. Pérez-Sáez, M. J., Montero, N., Redondo-Pachón, D., Crespo, M. & Pascual, J. Strategies for an expanded use of kidneys from elderly donors. Transplantation 101, 727–745 (2017).
22. Azancot, M. A. et al. The reproducibility and predictive value on outcome of renal biopsies from expanded criteria donors. Kidney Int. 85, 1161–1168 (2014).
23. Yin, P.-N. et al. Histopathological distinction of non-invasive and invasive bladder cancers using machine learning approaches. BMC Med. Inform. Decis. Mak. 20, 162 (2020).